
You need to cite the PAMI 2010 paper in all publications which describe work for which you used this software..
export PATH=$PATH:$HOME/mrf-src/tools
Then, build using the Makefile in the src directory:
cd mrf-src cd src make
The executable "mrf" can be found in "bin"
Note: the make process uses the makedefault software, which is integrated into the MRF package (no separate download is necessary).
The executable gives help when launched with the -h option:
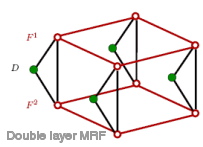
usage: /home/chris/bin/mrf [ options ]Options: [-P model-number] Choose the prior MRF model (def.: 1) 0 single MRF (autologistic or MLL) 1 double MRF (2x MLL / 2x Potts) 2 double MRF w line process 3 single MRF (Potts) [-O model-numer] Choose the obervation model (def.: 0) 0 double field - Gaussian grayvalue 1 double field - Gaussian color 2 double field - Generalized Gaussian grayvalue (only if numerical recipes are available) 3 single field - Gaussian grayvalue 4 double field - non-parametric 5 single field - Gauss corrected Sauvola [-I iterations ] iterations ICE (def.: 1) [-i iterations ] expansion move: n.o. cycles (def.: 10) simulated annealing: n.o. iter. gibbs sampler (def.: 300) [-h x,x,x,...] force values for the MRF parameters [-a adjust] adj. factor for beta parameter (graph cut only) (def.: 1) [-g iterations ] iterations ICM (greedy mode) (def.: 10) [-t t1 ] start temperature simulated annealing (def.: 2) [-c c ] parameter c simulated annealing (def.: 0.97) [-Z o.algo] Chose the optimization algorithm (def.: sima) sima simulated annealing and/or ICM grct graph cut [-B] Bleedthrough: separate result images into recto and verso part, even if a single MRF model is chosen. [-b] do use blurring in the model (not all models) [-D] use Derin et al. for a subset of parameters only [-M] do median filter before least squares estimation [-G] do Gauss filter before k-means [-s] do Sauvola et al.'s binarization instead of k-means (2 class segmentation, Potts model only). [-m] use the background substraction mask [-o color-code] color space and distance for kmeans init. (def.: 0) (0=rgb, 1=hsv, 2=L*u*v*, 3=L*a*b*, 4=L*a*b*+CIE94) [-C n.o.cl] number of classes (single MRF only) (def.: 3) [-V factor] constraint parm. for verso px (double MRF) (def.: -0.5) [-R] use gray.val. for the recognition of the recto class [-l] write the label field into the file 'labels.pgm' [-d option] write intermediate debug images to disk or not: 0: do not write 1: write visible segmentation output 2: write label images 3: write restored images Please specify an input and 3 output filenames!
The most important options are -P, which chooses the prior (classical single MRF or the proposed dobule MRF model), and -O which chooses the observation model (color or grayscale). The restorations in the PAMI paper have been obtained with the following parameters:
mrf -V -0.4 -Z grct -M -R -l -P 1 -O 0 inputimage output1 output2 output3
The only supported image format is Netpbm (.pgm and .ppm images). You can used the convert tool from the ImageMagick package to convert images to and from this format.
| File | Download | Remark |
| mrf | http | A binary compiled under Ubuntu 11.04 |
| src-mrf.zip | http | Linux sources |
This code has been written under and for Linux (Ubuntu 11.04). You also need the packages pdksh and ruby installed.
This program is free software; you can redistribute it and/or modify it under the terms of the GNU General Public License as published by the Free Software Foundation; either version 2 of the License, or (at your option) any later version.
This program is distributed in the hope that it will be useful, but WITHOUT ANY WARRANTY; without even the implied warranty of MERCHANTABILITY or FITNESS FOR A PARTICULAR PURPOSE. See the GNU General Public License for more details.
You should have received a copy of the GNU General Public License along with this program; if not, write to the Free Software Foundation, Inc., 59 Temple Place - Suite 330, Boston, MA 02111-1307, USA.
This software was written by Christian Wolf. Send comments, suggestions and bug reports to the follwing address:
christian.wolf (at) liris.cnrs.fr
You need to cite the PAMI 2010 paper in all publications which describe work for which you used this software..